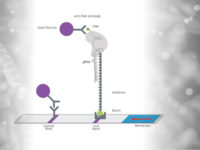
– Multiplexing is the ultimate challenge in the field of modern diagnostics. It’s much more than just the methodological “icing on the cake” – Techniques that allow strong multiplexing without significant loss of sensitivity & specificity will be “game-changers” for a new generation of field-applicable diagnostics –

The combination of nucleic acid analysis and lateral flow technology is without a doubt an important tool for current and future point-of-care and point-of-need applications. It doesn’t take a lot of imagination to assess the potential of these techniques. In most applications, the focus is on its simplicity, speed and robustness, whereby the performance of the analysis can be comparable to established laboratory reference methods. These characteristics qualify such tests for applications “in the field”. Terms such as specificity, robustness and sensitivity are mentioned again and again. But an important point undisputedly belongs to this list of “test-features”. We are talking about the generation of multiple information per analysis, which is called “Multiplexing”.
Multiplex Applications & the Universal Lateral Flow Platform – Milenia HybriDetect
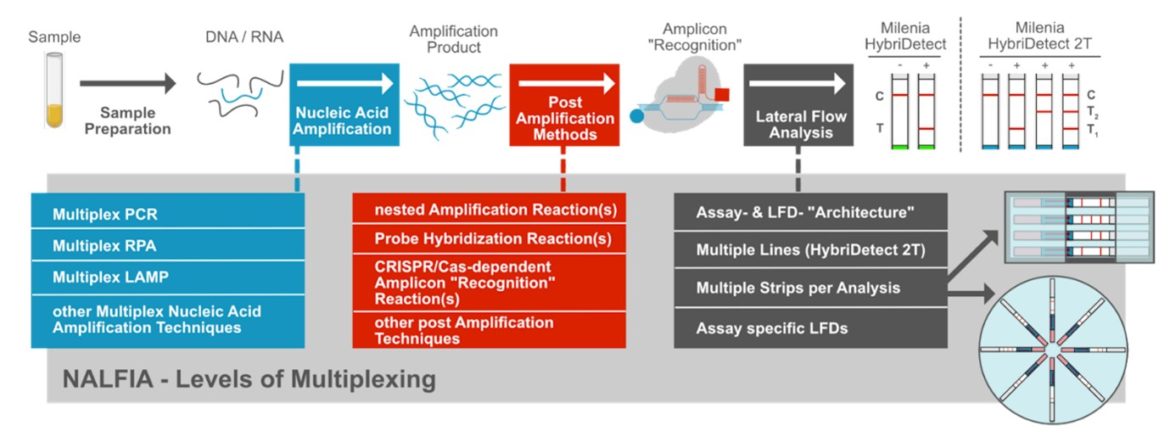
Milenia HybriDetect is a universal lateral flow development platform that allows scientists around the world simple rapid test development. The Milenia HybriDetect family includes two test strips, the Milenia HybriDetect and the Milenia HybriDetect 2T, whereby the 2T strip has 2 test lines and thus allows duplex applications. The multiplex options seem limited, especially for universal test strip formats, but there is actually still a lot of potential hidden in multiplexing in combination with the use of the universal lateral flow development platform – Milenia HybriDetect. The following figure describes that a certain degree of multiplexing can be generated at different points within a NALFIA (Nucleic Acild Lateral Flow Immunoassay. These “levels of multiplexing” must be perfectly coordinated with one another. The structure shown here also serves as a basis for discussion for this article, which is intended to provide an overview of existing technologies combining NALFIAs with multiplexing and at the same time facilitate the development of innovative test formats.
Learn more about the Milenia HybriDetect – General Detection Strategies
Learn more about the Milenia HybriDetect – Nucleic Acid Amplification & Lateral Flow
Advantage – Multiplex PCR & Lateral Flow
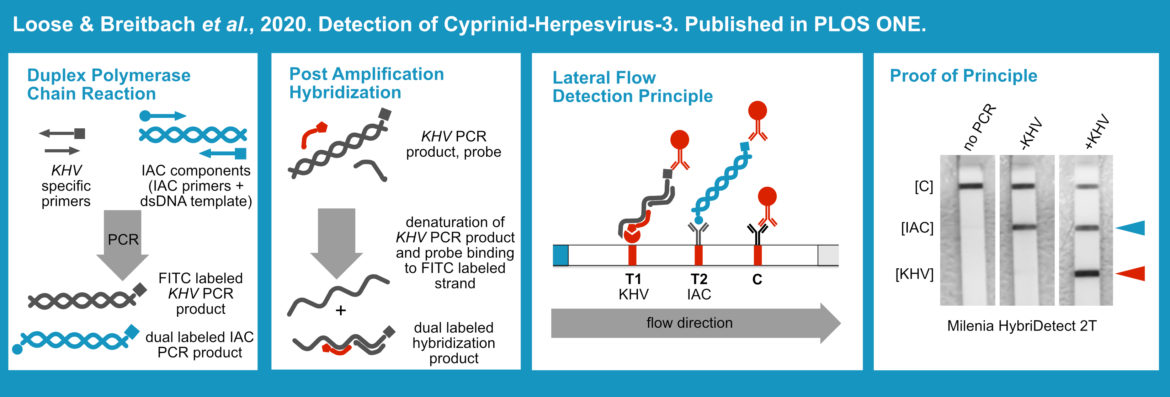
The PCR (Polymerase Chain Reaction) is the perfect choice, when it comes to multiplexing. Due to the simple primer design, simple adaptation of temperature protocols and the wide variety of optimized enzyme-systems, the PCR generally allows rapid development of well performing multiplex assays. Although PCR is discussed controversially in terms of Point-of-Care usability, this DNA-amplification technique has a lot of untapped potential in the context of NALFIAs. A few selected publications indicate the power of this technology as a basis for further multiplex applications & techniques. These selected publications use the universal Lateral Flow Device – Milenia HybriDetect 2T. [1] [5]
No DNA-Extraction? – DirectPCR and the Importance of an Internal Amplification Control
A wide variety of sample matrices can be used in PCR, if a suitable and robust enzyme system has been selected. This is the prerequisite for the direct application of a biological material in a PCR, eliminating the demand for a DNA or RNA extraction prior to the PCR reaction. This is an important feature for the development of point-of-care compatible rapid tests. Direct analysis from blood is just one example of practical relevance. The direct PCR format can benefit from the multiplex ability by implementing an IAC (Internal Amplification Control. This methodological approach allows a clear differentiation between negative results and matrix-dependent amplification inhibition. The Milenia HybriDetect 2T allows rapid evaluation of a direct-duplex PCR. [1] [5]
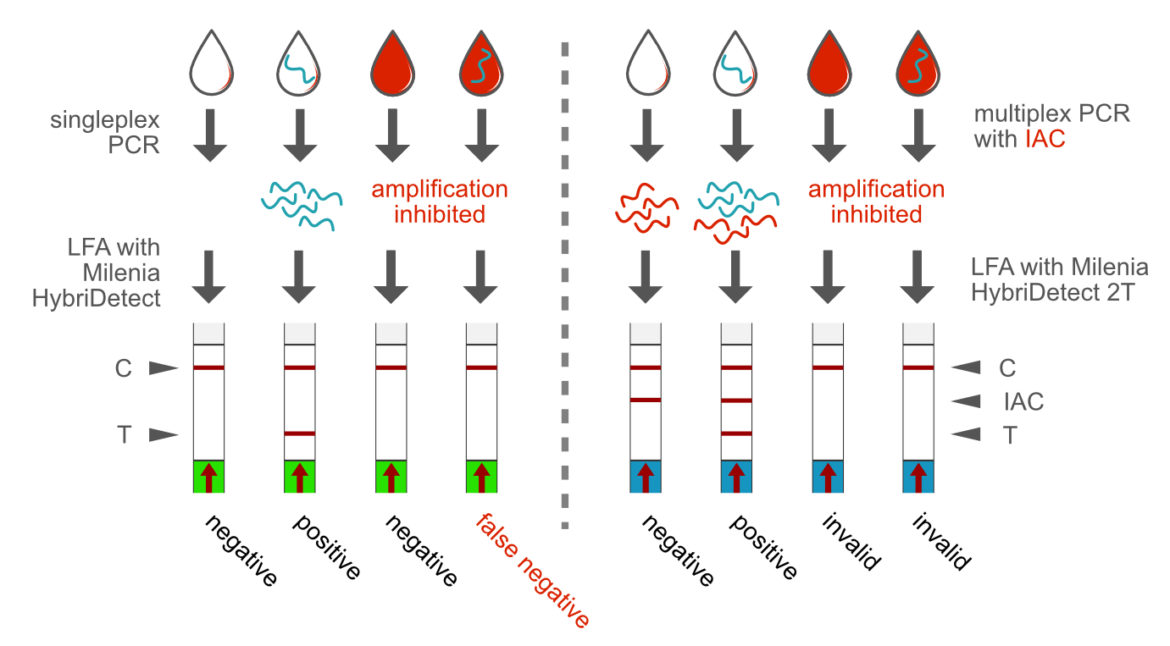
We at Milenia Biotec trust in this kind of technology. Therefore, we developed our own PCR-Lateral Flow-Kits including an extremely robust Polymerase that allows direct analysis without DNA-Extraction. Moreover, we developed an optimized IAC as perfect “Inhibition-Indicator” and of course, we use our own universal Lateral Flow Device, Milenia HybriDetect 2T, in order to specifically detect spoiling bacteria in beer and juice.
„Benefit from our experience, knowledge and products. Use the Milenia HybriDetect 2T for rapid development of duplex PCR applications. Ininitial development of personalized PCR-LFAs is possible within a few days to weeks.“
More Information about the Milenia GenLine Kits for rapid detection of Beer Spoiling Bacteria
More Information about the Milenia GenLine Kits for rapid detection of Alicyclobacillus spp.
Learn more about PCR & Lateral Flow
Multiplex RPA & Lateral Flow
Generally Recombinase Polymerase Amplification (RPA) is an isothermal DNA-amplification technique that allows extremely rapid and robust analysis. RPA is highly compatible with a simple-to-handle and -interpret Lateral Flow Readout, but it is not known to be the perfect technique for multiplexing. Nevertheless, it is possible to design duplex RPAs in order to generate two amplicons during the amplification reaction. Especially for this type of application an optimized Lateral Flow Assay Buffer was designed in order to suppress the disruptive character of RPA reagents against LFD-antibodies. This makes the Milenia HybriDetect 2T the perfect tool for analyzing multiplex RPA products. And this is exactly what has been proven in several scientific publications, which are listed below. [2] [6]
Two Informations for singleplex Quantification: Semi-quantitative RPA
Authors from Havard Medical School in Boston became creative with the presentation of an innovative Recombinase Polymerase Amplification (RPA) Assay design. They saw a Limitation in existing rapid POC assays because many of them rely on rapid DNA amplification methods, which amplify target-DNA extremely fast until saturation effects occur. In consequence there is no chance to (semi-) quantify the test outcome. In most cases it is a qualitative “yes” or “no” interpretation. Therefore, a competitive type of RPA was designed. Two differently labeled products are amplified at the same time in the same reaction tube. Amplicons are analyzed with a rapid lateral flow readout, using the Milenia HybriDetect2T. Combined signal intensities of the reference test line and the target test line allow semi-quantitative analysis. The more target DNA or RNA is introduced into the RPA reaction, the stronger the target test line appears on the LFD. Moreover, the intensity of the reference test-line decreases, because the “reference-amplicon” is less efficiently amplified. In contrast, if the amount of template decreases, test line intensity decreases as well and the intensity of the reference test line increases. This methodological approach allows a clear distinction between variable template quantities due to the design of a simple but very clever multiplex RPA variation. [7]
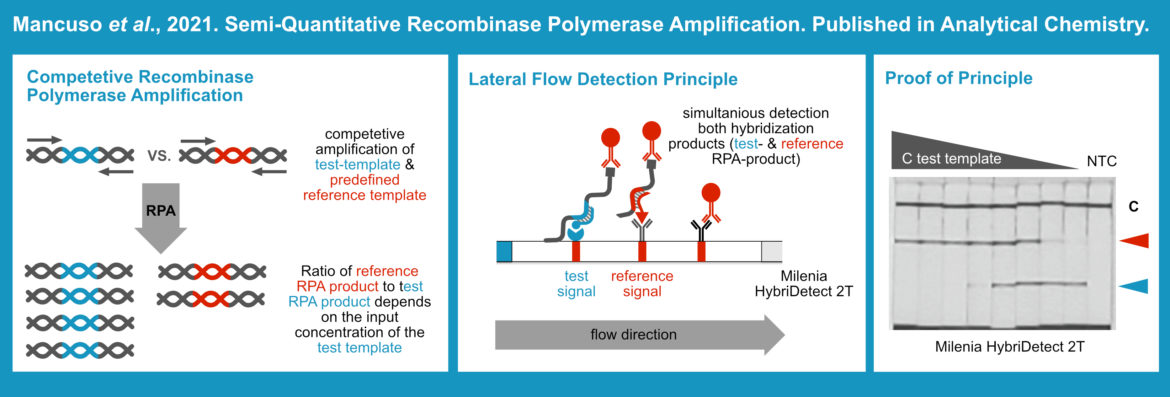
Link to the original publication by Mancuso et al.
Learn more about RPA & Lateral Flow
Multiplex LAMP
An efficiently performing singleplex LAMP-Assay needs 4 to 6 modular designed primers adressing 6 to 8 sequence-regions with defined properties & distances. This fact illustrates the complexity of the development of a well performing multiplex LAMP. But once a suitable primer combination has been found, the Loop mediated isothermal amplification (LAMP) can be an extremely impressive DNA amplification technique that is fast, highly sensitive, even in a multiplex setup. Beside standard duplex or triplex LAMP, there are more possibilities to achieve multiplexing, even with a singleplex Lateral Flow Device.
Two Amplification Products and One Signal – the High-Surety LAMP
Researchers from the University of Austin (Texas, USA) describe multiplexing as an important tool for the reduction of false positive and false negative results. The LAMP, an ideal low-tech DNA-amplification method for rapid, field-deployable testing, was multiplexed in order to detect SARS-CoV-2, the causative agent of COVID-19. Two established singleplex LAMP Assays were multiplexed, which worked out well. Moreover, the researchers developed amplicon-specific probes that were able to connect the LAMP-products of both reactions. Due to the chosen labeling strategy, only “connected” LAMP-products were detectable with the Milenia HybriDetect on one single test line. As a consequence, the presence of two different templates is necessary to generate a positive test result. The presence of just one amplifiable template allows amplification, but the Lateral Flow Device will give a negative result. The combination of a duplex LAMP-Assay and Lateral Flow Readout allowed sensitive detection of the SARS-CoV-2 with a significantly improved specificity. [3]
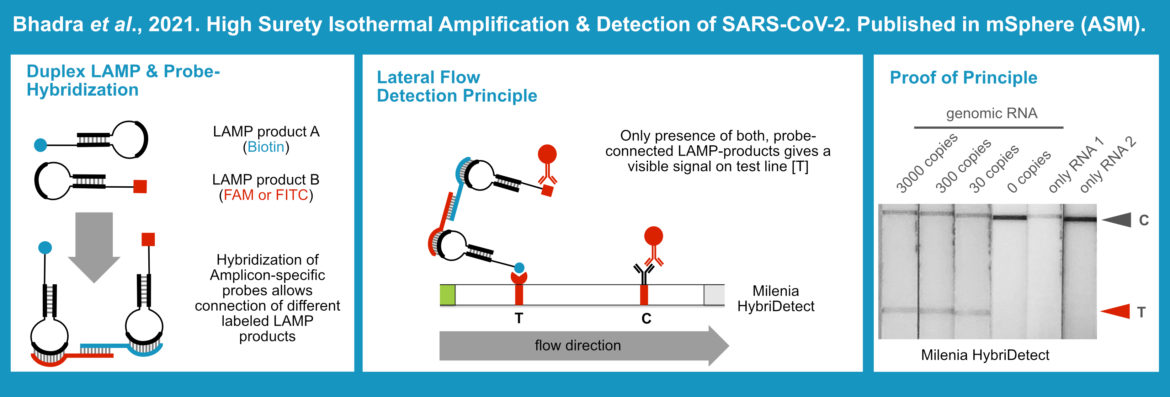
Link to the original publication by Bhadra et al.
Learn more about LAMP & Lateral Flow
The Addition of Post-Amplification Methods for stronger Multiplexing
Obviously, commercially available, universal Lateral Flow Devices are at some point limited when it comes to multiplexing. Nevertheless, there are ways to achieve a higher level of multiplexing by using additional techniques for amplicon “recognition” in combination with the use of multiple LFDs, in order to generate more information per analysis. This strategic approach has been described in several peer reviewed publications. The following sections describe three general post-amplification methods that may be useful for multiplexing: simple probe-Hybridization, CRISPR/Cas-dependent amplicon-”recognition” & nested amplification.
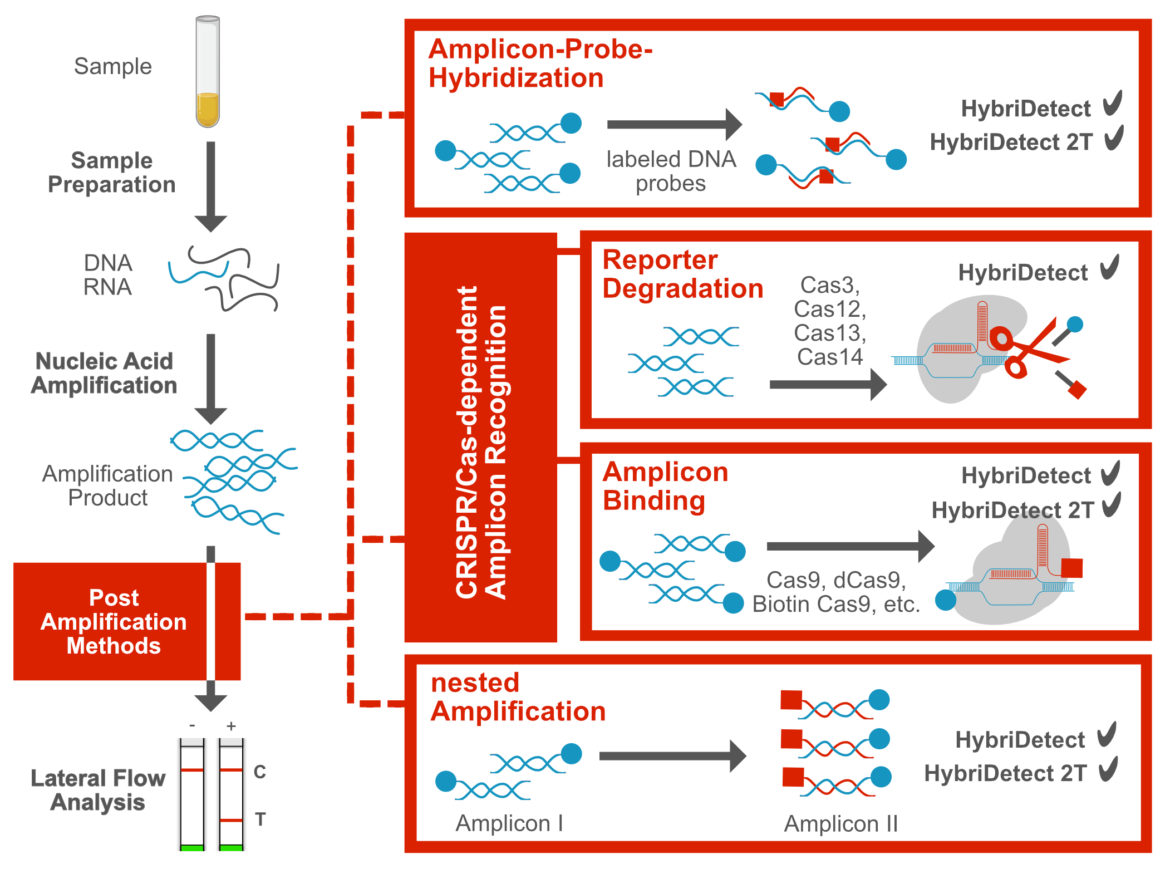
Amplicon-Probe-Hybridization
A very simple approach is the development of an additional hybridization reaction following the nucleic acid amplification reaction. For each hybridization reaction, 1 or 2 specific probes can be used to “recognize” amplicon-specific sequences. By establishing multiple hybridization reactions and using the universal test strips of the Milenia HybriDetect Platform, multiple pieces of information can be generated in one workflow. In internal experiments, hybridization in combination with various amplification techniques turned out to be a surprisingly powerful methodological tool, despite its simplicity. From genotyping, discrimination of SNPs to quite complex species differentiation, the simple hybridization of a labeled probe revealed an unexpected diversity. [1] [3] [6]
CRISPR/Cas dependent amplicon recognition
Over the last few years the CRISPR/Cas-”machinery” turned out to be an extremely valuable and versatile tool for modern molecular diagnostics. Detection sensitivities can be significantly improved, so that ultra-sensitive nucleic acid analyses are possible in a point-of-care or point-of-need setting. In addition, these molecular tools also allow an impressive level of selectivity. As a consequence, diagnostic CRISPR/Cas-dependent methods can achieve outstanding specificity. These properties make this technology a perfect post-amplification method that, with a little skill and creativity, can answer even complex questions in field-applications. A clear distinction has to be made between various CRISPR-Cas-related detection techniques. The best-known genome editing tool, CRISPR-Cas9, is used in the lateral flow context for applications in which gRNA-Cas9-amplicon-complexes are detected on the test of LFD. This principle (label incorporation strategy) is very suitable for multiplexing. In contrast, with Cas proteins such as Cas12 or Cas13 with high turnover (collateral nuclease activity), the status of a so-called dual-labeled reporter is analyzed. This enables very sensitive analyzes, which are less suitable for multiplex applications (label separation strategy). The following figure differentiates between these general methodological approaches.
Learn more about CRISPR & Lateral Flow
Learn more about general CRISPR-based diagnostics (Review from leading scientists in the field)
An Example from the Literature: CRISPR/Cas9 – Multiplex NALFIA
A very interesting publication from 2021 by Manoj Kumar, Sneha Gulati and colleagues illustrates the use of a SARS-CoV-2-specific amplification reaction, which is combined with CRISPR-Cas-dependent amplicon recognition assays for the differentiation of relevant SARS-CoV-2 variants. This is achieved by using several SNP-specific, FnCas9-mediated amplicon binding assays in combination with a simple lateral flow readout (Milenia HybriDetect). The sum of the results allows conclusions about the present SARS-CoV-2 variant. The authors called their method RAY which stands for Rapid variant AssaY. [4]
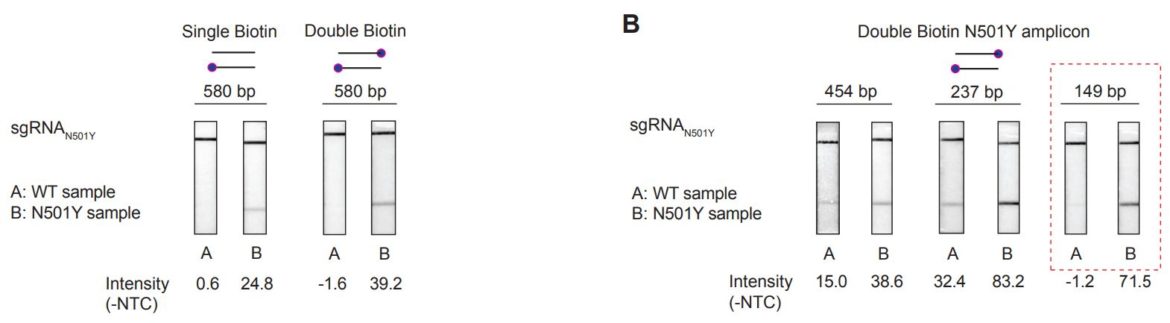
RAY for differentiation of SARS-CoV-2 Variants of Concern.
Request a detailled protocol for RAY Detection on Bio-Protocols.org
Nested Nucleic Acid Amplification
Nested amplification is a two-stage DNA amplification strategy in which the product of the first reaction serves as a template for the second one. In the follow-up reaction(s) several pieces of information can be generated. However, this form of additional analysis does not sound very attractive at first, because connecting a second nucleic acid amplification reaction to a first can be time-consuming, expensive and impractical. Nevertheless we see potential for innovation in this type of technology. The primer extension PCR (PEXT-PCR) can serve as an interesting example. Here, in a subsequent amplification, SNP-specific primer(s) are elongated in the 2nd PCR under strictly defined conditions, which result in specific reaction products that are detectable via Lateral Flow. By combining successive amplification reactions. Very sensitive and precise multiplex-test systems can be developed.
Collection of Relevant Multiplex NALFIA Publications
| Multiplex Detection of … | NALFIA Type | Reference |
|---|---|---|
| Cyprinid-Herpesvirus-3 & IAC | Duplex PCR & Hybridization | Loose & Breitbach et al., 2020 (1) |
| Cyprinid-Herpesvirus-3 & Carp Edema Virus | Duplex RPA (nfo-dependent) | Soliman & El-Matbouli, 2018 (2) |
| Two Targets SARS-CoV-2 | Duplex LAMP & Hybridization | Bhadra et al., 2021 (3) |
| SARS-CoV-2 Variants | PCR & CRISPR/Cas-Assay | Kumar et al., 2021 (4) |
| Authentification Cordyceps & IAC | Duplex PCR | Wong et al., 2015 (5) |
| Plasmodium Genus & P. knowlesi | Duplex RPA (nfo-dependent) | Lai et al., 2018 (6) |
| HIV / SARS-CoV-2 (semi-quantitative) | Competetive RPA & Hybridization | Mancuso et al., 2021 (7) |

- a b c : Loose, F. N., Breitbach, A., Bertalan, I., Rüster, D., Truyen, U., & Speck, S. (2020). Diagnostic validation of a rapid and field-applicable PCR-lateral flow test system for point-of-care detection of cyprinid herpesvirus 3 (CyHV-3). PloS one, 15(10), e0241420. https://doi.org/10.1371/journal.pone.0241420 ()
- : Soliman, H., & El-Matbouli, M. (2018). Rapid detection and differentiation of carp oedema virus and cyprinid herpes virus-3 in koi and common carp. Journal of fish diseases, 41(5), 761–772. https://doi.org/10.1111/jfd.12774 ()
- a b : Bhadra, S., Riedel, T. E., Lakhotia, S., Tran, N. D., & Ellington, A. D. (2021). High-Surety Isothermal Amplification and Detection of SARS-CoV-2. mSphere, 6(3), e00911-20. https://doi.org/10.1128/mSphere.00911-20 ()
- : Kumar, M., Gulati, S., Ansari, A. H., Phutela, R., Acharya, S., Azhar, M., Murthy, J., Kathpalia, P., Kanakan, A., Maurya, R., Vasudevan, J. S., S, A., Pandey, R., Maiti, S., & Chakraborty, D. (2021). FnCas9-based CRISPR diagnostic for rapid and accurate detection of major SARS-CoV-2 variants on a paper strip. eLife, 10, e67130. https://doi.org/10.7554/eLife.67130 ()
- a b : Wong, Y. L., Wong, K. L., & Shaw, P. C. (2015). Rapid authentication of Cordyceps by lateral flow dipstick. Journal of pharmaceutical and biomedical analysis, 111, 306–310. https://doi.org/10.1016/j.jpba.2015.04.003 ()
- a b : Lai, M. Y., Ooi, C. H., & Lau, Y. L. (2018). Recombinase Polymerase Amplification Combined with a Lateral Flow Strip for the Detection of Plasmodium knowlesi. The American journal of tropical medicine and hygiene, 98(3), 700–703. https://doi.org/10.4269/ajtmh.17-0738 ()
- : Mancuso, C. P., Lu, Z. X., Qian, J., Boswell, S. A., & Springer, M. (2021). A Semi-Quantitative Isothermal Diagnostic Assay Utilizing Competitive Amplification. Analytical chemistry, 93(27), 9541–9548. https://doi.org/10.1021/acs.analchem.1c01576 ()